scream() ensures that the structure of data is the same as
prototype, ptype. Under the hood, vctrs::vec_cast() is used, which
casts each column of data to the same type as the corresponding
column in ptype.
This casting enforces a number of important structural checks, including but not limited to:
Data Classes - Checks that the class of each column in
datais the same as the corresponding column inptype.Novel Levels - Checks that the factor columns in
datadon't have any new levels when compared with theptypecolumns. If there are new levels, a warning is issued and they are coerced toNA. This check is optional, and can be turned off withallow_novel_levels = TRUE.Level Recovery - Checks that the factor columns in
dataaren't missing any factor levels when compared with theptypecolumns. If there are missing levels, then they are restored.
Arguments
- data
A data frame containing the new data to check the structure of.
- ptype
A data frame prototype to cast
datato. This is commonly a 0-row slice of the training set.- allow_novel_levels
Should novel factor levels in
databe allowed? The safest approach is the default, which throws a warning when novel levels are found, and coerces them toNAvalues. Setting this argument toTRUEwill ignore all novel levels. This argument does not apply to ordered factors. Novel levels are not allowed in ordered factors because the level ordering is a critical part of the type.- ...
These dots are for future extensions and must be empty.
- call
The call used for errors and warnings.
Value
A tibble containing the required columns after any required structural modifications have been made.
Details
scream() is called by forge() after shrink() but before the
actual processing is done. Generally, you don't need to call scream()
directly, as forge() will do it for you.
If scream() is used as a standalone function, it is good practice to call
shrink() right before it as there are no checks in scream() that ensure
that all of the required column names actually exist in data. Those
checks exist in shrink().
Factor Levels
scream() tries to be helpful by recovering missing factor levels and
warning about novel levels. The following graphic outlines how scream()
handles factor levels when coercing from a column in data to a
column in ptype.
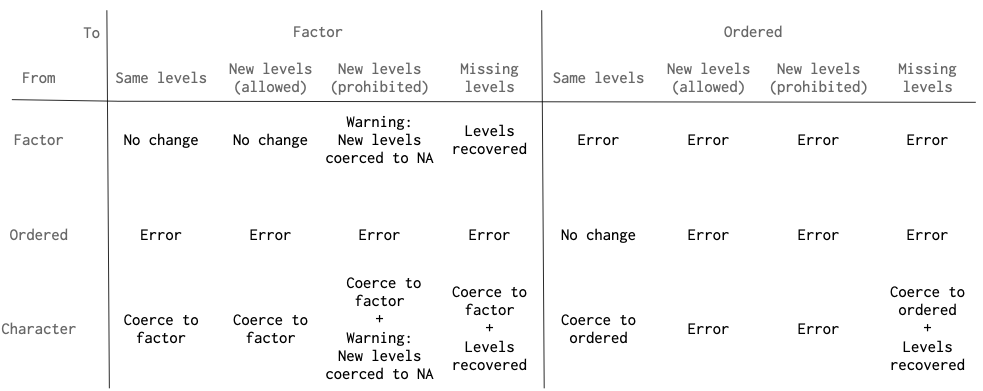
Note that ordered factor handing is much stricter than factor handling.
Ordered factors in data must have exactly the same levels as ordered
factors in ptype.
Examples
# ---------------------------------------------------------------------------
# Setup
train <- iris[1:100, ]
test <- iris[101:150, ]
# mold() is run at model fit time
# and a formula preprocessing blueprint is recorded
x <- mold(log(Sepal.Width) ~ Species, train)
# Inside the result of mold() are the prototype tibbles
# for the predictors and the outcomes
ptype_pred <- x$blueprint$ptypes$predictors
ptype_out <- x$blueprint$ptypes$outcomes
# ---------------------------------------------------------------------------
# shrink() / scream()
# Pass the test data, along with a prototype, to
# shrink() to extract the prototype columns
test_shrunk <- shrink(test, ptype_pred)
# Now pass that to scream() to perform validation checks
# If no warnings / errors are thrown, the checks were
# successful!
scream(test_shrunk, ptype_pred)
#> # A tibble: 50 × 1
#> Species
#> <fct>
#> 1 virginica
#> 2 virginica
#> 3 virginica
#> 4 virginica
#> 5 virginica
#> 6 virginica
#> 7 virginica
#> 8 virginica
#> 9 virginica
#> 10 virginica
#> # ℹ 40 more rows
# ---------------------------------------------------------------------------
# Outcomes
# To also extract the outcomes, use the outcome prototype
test_outcome <- shrink(test, ptype_out)
scream(test_outcome, ptype_out)
#> # A tibble: 50 × 1
#> Sepal.Width
#> <dbl>
#> 1 3.3
#> 2 2.7
#> 3 3
#> 4 2.9
#> 5 3
#> 6 3
#> 7 2.5
#> 8 2.9
#> 9 2.5
#> 10 3.6
#> # ℹ 40 more rows
# ---------------------------------------------------------------------------
# Casting
# scream() uses vctrs::vec_cast() to intelligently convert
# new data to the prototype automatically. This means
# it can automatically perform certain conversions, like
# coercing character columns to factors.
test2 <- test
test2$Species <- as.character(test2$Species)
test2_shrunk <- shrink(test2, ptype_pred)
scream(test2_shrunk, ptype_pred)
#> # A tibble: 50 × 1
#> Species
#> <fct>
#> 1 virginica
#> 2 virginica
#> 3 virginica
#> 4 virginica
#> 5 virginica
#> 6 virginica
#> 7 virginica
#> 8 virginica
#> 9 virginica
#> 10 virginica
#> # ℹ 40 more rows
# It can also recover missing factor levels.
# For example, it is plausible that the test data only had the
# "virginica" level
test3 <- test
test3$Species <- factor(test3$Species, levels = "virginica")
test3_shrunk <- shrink(test3, ptype_pred)
test3_fixed <- scream(test3_shrunk, ptype_pred)
# scream() recovered the missing levels
levels(test3_fixed$Species)
#> [1] "setosa" "versicolor" "virginica"
# ---------------------------------------------------------------------------
# Novel levels
# When novel levels with any data are present in `data`, the default
# is to coerce them to `NA` values with a warning.
test4 <- test
test4$Species <- as.character(test4$Species)
test4$Species[1] <- "new_level"
test4$Species <- factor(
test4$Species,
levels = c(levels(test$Species), "new_level")
)
test4 <- shrink(test4, ptype_pred)
# Warning is thrown
test4_removed <- scream(test4, ptype_pred)
#> Warning: Novel level found in column "Species": "new_level".
#> ℹ The level has been removed, and values have been coerced to <NA>.
# Novel level is removed
levels(test4_removed$Species)
#> [1] "setosa" "versicolor" "virginica"
# No warning is thrown
test4_kept <- scream(test4, ptype_pred, allow_novel_levels = TRUE)
# Novel level is kept
levels(test4_kept$Species)
#> [1] "setosa" "versicolor" "virginica" "new_level"
